Democratizing Discovery: Large Language Models (LLMs), Hackathons, and the Future of Materials Science and Chemistry Research
Or: How To Get More Scientists to Build with AI Agents
Large language models (LLMs) are rapidly changing scientific research. Anthropic, OpenAI, Google, FutureHouse, and others have all shared recent work documenting the expansive scope. The claims range from e.g., compressing six person-months of research into a day; modeling how small molecules interact with proteins with dramatically increased speed and precision; and solving long-standing mathematical conjectures. Beyond headline-grabbing scientific results LLMs are reshaping the day-to-day work with Novo Nordisk reporting usage of Claude to reduce paperwork overhead dramatically. This is happening now, and if the models and tooling keep improving it will only accelerate.
Research in materials science and chemistry presents a unique opportunity with specific bottlenecks that align with what LLMs are unusually well-suited to address. The friction points are addressable, and the vision of what is possible on the other side: better batteries, new classes of drugs, lighter structural materials, more efficient catalysts, is deeply important to the future of material abundance, economic prosperity, and good health that we hope for.
The Bottlenecks are Specific and Addressable
Work in materials science and chemistry spans extraordinary ranges in length scales (from a few atoms to manufactured components), times scales (from femtosecond reaction dynamics to years for material degradation), and methodologies (from quantum chemistry simulations to bench-scale synthesis and production scale up). The result is exceptional heterogeneity in data, tools, and workflows. A team optimizing a catalyst might need to combine campaigns of quantum simulations and manage data from dozens of instruments with differing file formats — all while synthesizing their own knowledge with figures, tables, and text from thousands of papers of varying quality.
Further, while materials science and chemistry have generated large datasets and repositories (e.g., Materials Project, OQMD, NOMAD, OMol25, the Materials Data Facility and more - see this list of 100s of other resources) there isn’t an equivalent to GenBank for genomics or Protein Databank for protein science because of the exceptional heterogeneity of materials and chemistry data. As such, there is no universal structured repository of synthesis procedures or process-structure-property relationships or molecular equivalent, and instead that information is often contained in personal expertise, millions of papers, figures, tables, hundreds of data resources, notebooks, and various repositories.
Researchers still spend significant time on mind-numbing tasks like data entry, extracting information from papers by hand, converting files between formats, and writing documentation and reports. They unfortunately must spend time on the small, grinding tasks that have nothing to do with the actual creative work of science but consume enormous chunks of mental capacity.
Dealing with such data and software heterogeneity and varying workflows are tasks that LLMs are particularly good at. LLMs have the potential to become connective tissue – a universal interface layer – because they are particularly adept at translating between heterogeneities in e.g., human intent/machine actions, multiple schemas, inputs/outputs across tools, and even narrative scientific context and structured data procedures. With such a broad new class of capabilities, there is a transformational opportunity for this connective tissue to be built between fragmented software, infrastructure, databases, papers, and more that currently don’t talk to each other. In such a vast application space of rapidly emerging LLM capabilities, it would take years to fully understand, specify, prioritize, and build such infrastructure if we waited for traditional research projects to be funded for this kind of work.
Instead, we decided to see what tools scientists could build for themselves through a hackathon.
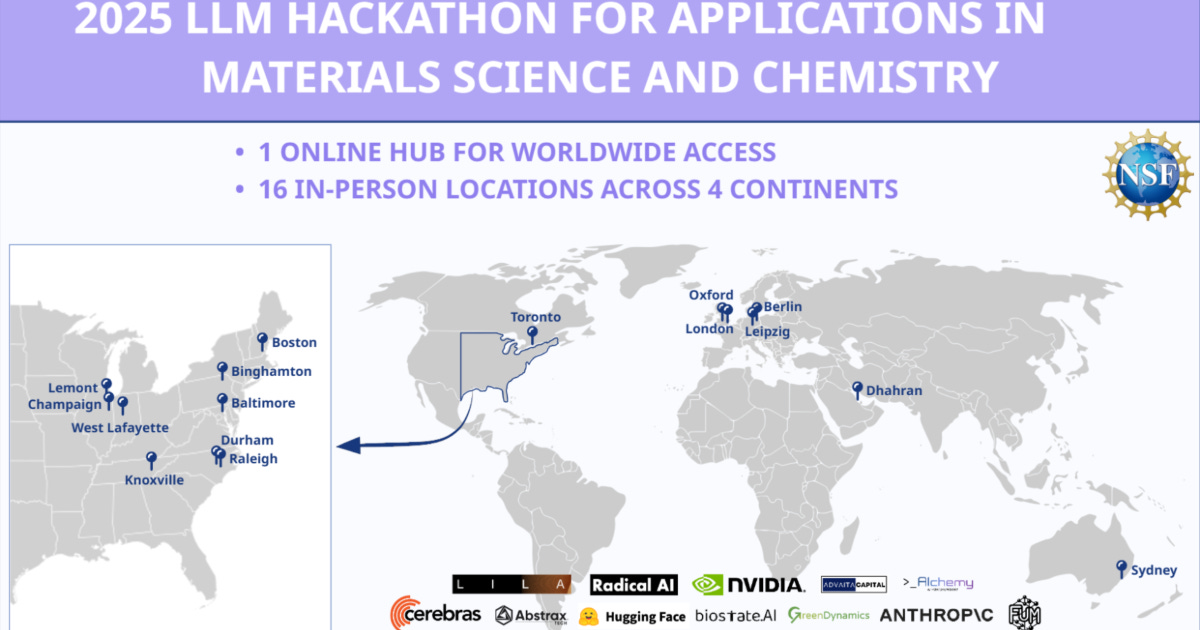
The Hackathons: 2000 researchers across 4 continents
The first LLM Hackathon for Applications in Materials Science and Chemistry was held in 2023, and for this event, we didn’t know if anyone would show up or if the tools were mature enough to build anything real in 24 hours. It also wasn’t assured that materials scientists and chemists would have the requisite software development skills. But people showed up, and the latent capability to build was evident almost immediately. Many had never built an agentic system or worked with an LLM before.
Over three events, in 2023 and subsequently 2024, and 2025 (see figure), more than 2,000 participants, primarily graduate students, created over 170 publicly documented, open-source projects. The 2025 event alone had 16 in-person sites, plus an online hub for worldwide access. We provide in detail, a description of all of the projects and links here, but in short, they built complex software including: natural language interfaces that let non-experts control advanced instruments; agentic workflows that compress the time from idea to experiment; automated data management systems that make it possible to share data across labs; new ways to train researchers; new generative and predictive models; tools to extract structured information from a corpus of research papers; and much more.
These events produced moments of connection, learning, and building. This included teams spanning the globe and formed in interesting ways, e.g., researchers in India partnering with researchers in the United States and midnight “aha” moments on Slack that put a team on a prize-winning path. We purposefully designed this not as only a single day event, but as an ongoing community where these moments have a space to continue, enabling researchers to catalyze startups and worldwide collaborations, land fellowships, write papers together, and make discoveries of their own.
In these events, we primarily focused on building and learning. To be clear, these projects are prototypes, not yet production systems. But many of them are really good prototypes, and the speed at which they were built tells us something important about where the technology is right now and the latent capability of researchers to adapt these tools.
We next describe the outcomes and what we learned in more detail. We group the projects into four buckets for convenience, though many projects span multiple categories including:
Prediction and design: using LLMs alongside traditional ML to e.g., predict material properties or generate candidate structures or other distributions, especially when data are scarce.
Interfaces and automation: conversational control of instruments, simulations, databases, and development of closed-loop systems that compress the hypothesis-to-validation cycle.
Data management and knowledge extraction: compiling structured, computable data out of the messy world of PDFs, lab notebooks, patents, lightly documented datasets, and inconsistent databases.
Education and scientific communication: tutoring platforms, virtual lab simulators, tools for creating explanatory content to connect with audiences ranging from aspiring students to domain experts.
Prediction and design: LLMs as complements, not replacements
In materials science and chemistry, researchers are often working in low data regimes and with heterogeneous data as discussed previously. Importantly, the projects showed the strength of LLMs to work under such conditions.
The LLM4ConProp team explored whether LLMs can directly predict a material property of concrete. They assembled two curated datasets totaling nearly 4,000 records of concrete compressive strength and material composition and processing, and ran a head-to-head comparison of GPT-4.1 in zero-shot and few-shot settings versus tree-based models (random forests, XGBoost, LightGBM). The results showed that with enough in-context examples, the LLM approached ML baselines for low data regimes.
Unsurprisingly, LLMs also work well with multimodal and heterogeneous inputs. Researchers may wish to provide natural language description of desired properties, leverage research papers, or describe material or molecular characteristics and generate structures, synthesis pathways, or other distributions. The MIDAS team built a prototype showing structure-based drug design by conditioning a diffusion model on both protein pocket geometry and natural language instructions. With this system, users can say things like “generate a molecule with a hydroxyl group targeting this binding site” and get candidates that reflect both the structural constraints and chemical intuition. To train this, the team generated approximately one million molecule-text description pairs using GPT-3.5, covering functional groups, molecular properties, and pharmacophore descriptions. The whole system was wrapped in a chat interface with additional tools for docking analysis, similarity search, retrosynthesis, and iterative refinement through conversational feedback. Another project, SKY, tackled the inverse problem of how to make a material given specific property targets. The system takes a natural language description of a target structure, runs recursive similarity searches across Materials Project data, and uses LLM reasoning to propose grounded synthesis routes.
These projects illustrate how LLMs are effective in prediction and design especially when paired with traditional ML models as complements and to provide interfaces that handle the messy, heterogeneous inputs, outputs from these models, and to handle cross-service data connections. As foundation models improve at reasoning over mixed modalities like text, structures, spectra, images, this complementary role is likely to expand, particularly in the low-data regimes that define much of materials science and chemistry.
Interfaces and automation: making billion-dollar infrastructure more accessible
The US spends billions annually on national user facilities - synchrotrons, neutron sources, nanocenters, high-end electron microscopes, exascale supercomputers - that are too large and specialized for any single university to host. These are some of the most powerful scientific instruments on the planet, including publicly funded supercomputing facilities that provide compute access to researchers nationwide. Far fewer researchers use them than should, largely because accessing them requires months of specialized training. Making these facilities accessible without months of training could have outsized impact.
Natural language interfaces offer a clear path forward. Some of the most common applications run on scientific supercomputing are density functional theory (DFT) calculations and molecular dynamics (MD), but running simulation campaigns at high-performance computing facilities requires understanding of the specific system you are running on, optimizing for scale across many nodes, carefully monitoring the jobs, and more. LARA-HPC built an assistant that translates scientific goals, e.g., “compute the atomization energy of HCN”, into complete HPC DFT workflows. Similarly MINT LLM created natural language interfaces for MD analysis across codes.
Automated labs are gaining increased prominence within national user facilities and industry. The ACME team connected molecular design, quantum chemistry, and simulated robotic experimentation in a single feedback loop for discovering molecules for critical materials extraction. A reasoning model retrieves domain knowledge, e.g., ligand design rules, coordination geometry constraints from curated publications, then constructs candidate molecules. Those candidates pass through automated computational screening where the system builds 3D structures, runs semi-empirical quantum chemistry (GFN-xTB), and ranks by metal-ion binding affinity. In this hackathon, the top candidates didn’t enter the physical world, but rather a virtual stand-in that is being developed to synthesize and characterize coatings of these future molecules. Simulation results feed back to the reasoning model, which updates design rules and proposes improved candidates.
These prototypes are directly aligned with major national investments. The US Genesis Mission, launched by executive order in late 2025, aims to connect DOE’s 17 national laboratories, user facilities and scientific datasets into an integrated discovery engine. Canada’s Acceleration Consortium, various efforts in Europe and Asia, and industry efforts from Lila, Cusp, DeepMind, Periodic Labs, Radical AI, and others are pursuing parallel ambitions spanning automated labs and scientific intelligence. The hackathon projects show that the research community is ready to build on this infrastructure, and that many of these ambitious goals may be closer than expected. These working prototypes also point to a future where scientific user facilities have new interfaces for improved accessibility and efficiency.
Data management and knowledge extraction: liberating a century of science from PDFs and notebooks
Federal mandates increasingly require that scientific data be Findable, Accessible, Interoperable, and Reusable. In practice, that compliance falls disproportionately on graduate students and postdocs, the benefit to any single researcher rarely feels worth the effort, and the result is repositories that are technically FAIR but still unusable. LLMs offer a way to fundamentally shift this burden.
LLMs offer a way to fundamentally shift this burden and capture structured data more efficiently. For example, ExpAlign created a pipeline that parses hundreds of research PDFs, extracts key properties and hidden experimental details, flags inconsistencies, and combines results into clean, ML-ready data tables. MuMMIE tackled multilingual patent extraction across five languages, creating benchmarks for cross-lingual scientific information extraction. SuperconLLM built a four-agent pipeline that screens arXiv papers, performs named entity recognition and relation extraction, and generates structured database entries for superconductor properties. PolyNexus created a domain-specific knowledge base for electroactive polymers where you can ask natural language queries (”What is the conductivity of PEDOT:PSS?”) and get citation-backed results.
This extraction and information access at scale, when coupled with private sector efforts (e.g., Edison Scientific/Future House, Chemical Abstracts Service, ChemDataExtractor, Citrine Informatics), promises to collate and liberate hundreds of years of humanity’s collective scientific knowledge from PDF files into something structured, computable, and accessible data sources.
Education and scientific communication: the parts nobody talks about
Research impact depends not just on discovery but on how effectively knowledge transfers to the next generation of researchers and diffuses to adjacent fields, and to the public. These are areas where researchers typically receive little training and have few tools, and where LLMs may have an outsized impact.
For example, ChemTutor AI built a domain-specific tutoring platform that generates personalized problems with interactive 3D molecular visualizations and adaptive difficulty. It provides pedagogical scaffolding including e.g., step-by-step reasoning, worked examples, and Socratic questioning. MatSci LapLab, built a tool to simulate characterization techniques like TGA, SEM, and tensile testing, enabling students to train virtually.
On the communication side, AtomicShorts built a three-agent pipeline that helps scientists create short explanatory videos at a fraction of commercial costs and at different audience knowledge levels. By keeping scientists in the loop, the system ensures accuracy while reducing the enormous gap between researchers and the public.
These tools address an asymmetry, i.e., researchers are trained and incentivized to do science but less so to teach or communicate efficiently. If LLMs can lower the cost of creating high-quality educational and explanatory content, the pool of people who engage with, and eventually contribute to, scientific research could increase significantly.
The hackathon model: why this worked and how to replicate it
The hackathon series was, itself, an experimental test of what we call the “Social-First Hackathon Model”. This model is one that many research communities could replicate, and in fact, has already been used for two hackathons in microscopy and one in Bayesian optimization.
We designed the model around five principles:
Public and social by default. Teams submit entries via social media, LinkedIn, X, YouTube, with accompanying code repositories and a Google Forms entry. The social posts create immediate public visibility, provide verifiable credit for CVs, and eliminate the need for custom infrastructure.
Minimal central coordination. The entire event runs on free tools like Slack, Zoom, GitHub, Luma, and Google Workspace. Participants form teams, decide topics, and plan projects autonomously. Two to three central organizers coordinated an event for over a thousand participants.
Hybrid from the start. A global virtual cohort participates alongside physical hubs at institutions that volunteer to host. This decentralizes operations while maintaining a unified program and allows the best talent from anywhere to participate.
Time-boxed intensity. The 24h constraint forces rapid prioritization, prevents scope creep, allows researchers to participate without compromising their ongoing research, and creates urgency.
Leaning into academic incentives. Participants are primarily graduate students and postdocs. Co-authorship on papers, presentation opportunities, awards for CVs matter. After each event, we assembled teams to write articles including as many active participants as possible.
Three lessons emerged across events. First, team formation before the event is critical. The most common failure mode is participants showing up without teammates and being unable to crystallize both a team and a concept. The 2025 event addressed this by emphasizing early skill-matching for over a month before the event, using the shared Slack, virtual meetings, and a custom Miro board.
Second, domain breadth within teams matters. The events have no limits on team formation. Teams can be as big or small as needed, and combine in-person and virtual. Many of the strongest teams brought together broad teams including e.g., computational chemists, experimentalists, and ML engineers.
Third, the community persists. Over 1,400 researchers remained active in shared Slack channels after the events concluded, continuing collaborations, posting job opportunities, and building on each other’s work. The hackathon is a nucleation point that brings together high-agency researchers in a long-lived community that continues to percolate afterward.
Importantly, this model handles aspects that traditional funding mechanisms handle poorly: e.g., 1) rapid landscape mapping (170 projects surveying possibilities), 2) real-time workforce development via participants learning by building and forming lasting collaborations, 3) public-by-default outputs where every project is immediately available for others to learn from and build on, 4) and talent identification where two days of effort can produce a significant addition to the researcher’s CV.
The projects and community described here focused on materials science and chemistry, but illustrate a broader pattern about how AI capabilities spread through research communities. Properly structured hackathons may function as adoption accelerators, with the ability to compress the learning curve, give researchers permission to experiment (and fail), and produce reusable examples and visible proof that the tools work.
These events showed not just the diffusion, but the building of capability, showing how a capable workforce, able to effectively leverage AI can be built rapidly. The hackathons were views into the diffusion and capacity building happening in real time, suggesting that research institutions, national labs, and professional societies could drive meaningful AI adoption and diffusion of capabilities and knowledge by running similar events in their own domains. The model is lightweight, replicable, and documented. However, there are still areas for improvement, e.g., sustaining momentum after the event ends, securing compute credits and inference access for deeper development, and bridging the gap between a promising hackathon prototype and a tool that researchers use daily. These are tractable problems, but they require intentional investment in the community infrastructure, scientific middleware, and shared compute to further build researcher capacity and enable diffusion of tools into production use.
One important point sticks with me and fills me with hope. Many of the teams that participated, including those that built reasoning agents connected to simulation tools and external databases, autonomous NMR analysis pipelines, and natural language interfaces for software and hardware had never worked with LLMs or constructed an agentic system before. Yet, with a clear goal, and aligned incentives, they accomplished it together in just over a day. That fact alone should change our priors on the near-term trajectory of scientific research, even if these participants are perhaps exceptional due to selection biases.
Scientific tools often diffuse slowly through research communities, often taking years between invention and widespread use. The 170 projects built at the events are prototypes that help provide a concrete starting point to speed the diffusion. Yet, there is real work between a hackathon demo and a tool used daily by thousands of researchers that requires additional work and investment. From these events, researchers have catalyzed new collaborations, presented results at top international conferences, secured new funding, created teaching modules, and created software used by many other research groups. All of this, paired with the scale and breadth of working demonstrations show how much is now possible promising to draw more people in to build the next set.
This diffusion also requires building researcher expertise and familiarity. The hackathon events showed that with even a modest incentive structure and a focused community-building effort, thousands of researchers with minimal initial LLM training can be trained via “learn-by-shipping”. In this model each team produces (1) a concept, (2) a software repository, (3) a demo artifact (e.g., video), and (4) a short explainer. Those outputs and the community itself act as a diffusion substrate creating code others can build upon, workflows others can copy, and explorations of spaces relevant to many different research groups. As such, the training does not stay local to participants. This creates a virtuous cycle where increased visibility recruits the next cohort, reuse of the examples turns prototypes into shared resources or production software, and where normalization lowers perceived risk and increases adoption inside labs.
As these tools propagate, and e.g., the ability to run and interpret common techniques like XRD analysis, DFT, MD, molecular dynamics or leverage unique user facilities becomes conversational and ubiquitous, the barriers between disciplines become more porous with AI tools reducing transaction costs, e.g., meaning less time negotiating data formats, learning one-off software stacks, or translating jargon. Further, as data management overhead disappears, researchers can reclaim time for actual science and other research teams get access to better open data to use in their analyses. Towards training the next generation using AI, students can train on simulated characterization techniques expanding the researcher capacity dramatically. Breakthrough science has always thrived at the intersections between fields, institutions, capabilities, and expertise. In many ways, these projects are helping to create new intersections. AI is set to change dramatically as user expertise in AI tools diffuses, models improve, creating more of those intersections.
None of these tools or approaches are guaranteed to become widely adopted. The new technology creates an opening, but realizing the benefits depends on whether the research community and the institutions and agencies that support the research ecosystem lean into the opportunity. The experiences documented here suggest that relatively modest investments in inference infrastructure (or cloud credits), scientific middleware, documentation standards, and community coordination could yield outsized returns.
If you are an interested scientist, you can join the 1400+ researchers continuing to build in this space on the hackathon slack. Reach out to me if you’d like to sponsor this work, visit our community, and stay tuned for our next events!